Hi,
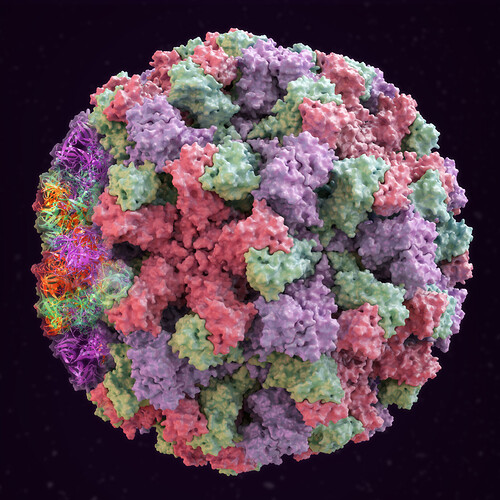
We’re a newly launched start-up doing biological simulations, in order to assess the efficacy of a novel class of drugs. We need an animation showing how a virus capsid assembles. I have already bought this static model of a norovirus from turbosquid:
https://www.turbosquid.com/3d-models/3d-norovirus-norwalk-virus-1354514
The model already has 60 sub-components (each subcomponent has 3 proteins each). We need to show how the proteins come together one-by-one to create the full virus particle. To achieve this, it’s easier to start from the full virus and simulate a dis-assembly => and then run the animation backwards. So you start with the full virus capsid, and take each protein one by one (60 in total) and start diffusing them through a random walk in the 3D space. Then you put together the movie in reverse, to show assembly!
Our model has two views: ribbon and surface.
The virus has two types of models: ribbon and surface. The animation will use the ribbon model throughout, and only overlay the surface at the end, for the final 5 seconds. The entire animation should be for 30-40 seconds.
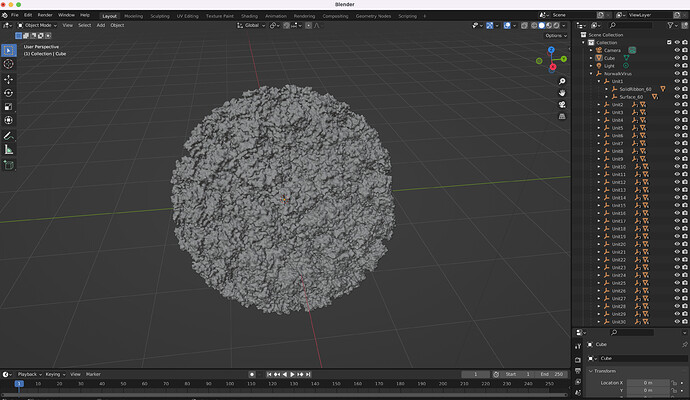
The model has 6 million polygons, and was natively done in Maya 2017, but we also have .FBX and .OBJ, so can be imported into Blender, 3ds Max, etc … Below is a screenshot of the model loaded in Blender, showing the 60 sub-components.
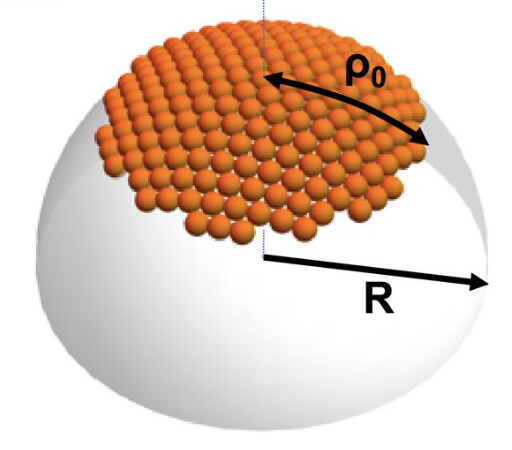
An important aspect of the animation is that the capsid should grow from a core nucleus, like in the image below. So new proteins are only added on the periphery of the partial capsule.
Let me know if you can help. I would really appreciate it. Many thanks!